Information and Semiosis in Living Systems: A Semiotic Approach
João Queiroza,b, Claus Emmechec and Charbel Niño El-Hania
© This paper may not be reproduced without the permission of the authors
ABSTRACT
During the 1950s and 1960s, genetics and cell and molecular biology have been swamped by terms borrowed from information theory. This ‘information talk’ still pervades these fields, including widely used terms such as ‘genetic code’, ‘messenger RNA’, ‘transcription’, ‘translation’, ‘transduction’, ‘genetic information’, ‘chemical signals’, ‘cell signaling’ etc. As the concept of information and its plethora of associated notions were introduced in biology, several problems emerged, with which the tradition of biology was unprepared to cope. Instead of deepening the discussion about ‘information talk’, the trend in the biological sciences was one of treating ‘information’ as merely sequence information in DNA or proteins. Today, a number of researchers consider information talk as inadequate and ‘just metaphorical’, expressing a skepticism about the use of the term ‘information’ and its derivatives in biology as a natural science. We disagree with this position, claiming instead that the notion of information and other related ideas grasp some fundamental features of biological systems and processes that might be otherwise neglected. Our problem is not to get rid of information talk, but rather to clarify it by using a proper theoretical framework. We intend to show that the use of semiotic concepts and theories to interpret information talk can contribute to the construction of a precise and coherent account of information in biology. For this purpose, we introduce here a model of information as semiosis, grounded on Peircean semiotics. Peirce’s formal science of signs provides an analytic framework in which information can be modeled as a pragmatic triadic dependent process that irreducibly connects signs, objects, and interpretants (effects on interpreters). According to the model developed in this paper, information is treated as semiosis, i.e., the communication of a form or habit from an object to an interpretant through a sign, so as to constrain (in general) the interpretant as a sign or (in biological systems) the interpreter’s behavior. We employ this treatment of information for building an account of genes as signs and genetic information as semiosis.
Keywords: Gene, Information, Signs, Semiosis, Biosemiotics, C. S. Peirce.
1. INTRODUCTION
‘Information’ is a concept which is very important but problematic in biology (see Oyama 2000, Stuart 1985, Sarkar 1996, Griffiths 2001, Jablonka 2002). The concept of information in biology has been recently a topic of substantial discussion (See, e.g., Maynard Smith 2000, Godfrey-Smith 2000, Sarkar 2000, Sterelny 2000, Wynnie 2000, Jablonka 2002, Adami 2004). Furthermore, the evolution of new kinds of information and information interpretation systems in living beings has received a great deal of attention recently (See, e.g., Jablonka 1994; Jablonka and Szathmáry 1995; Maynard Smith and Szathmáry 1995, 1999; Jablonka, Lamb & Avital 1998). It is even the case that the evolution of different ways of storing, transmitting, and interpreting ‘information’ can be treated as a major theme in the history of life (Maynard Smith and Szathmáry 1995, 1999; Jablonka 2002).
As Griffiths (2001) wrote recently, ‘genetic information’ is a metaphor in search of a theory. We think the same can be said, in general terms, of the current use of the term ‘information’ and its derivatives in biology. After all, a number of researchers consider all information-talk as inadequate, taking a skeptical view towards the very use of the term ‘information’ and its derivatives in biology as a natural science (e.g., Stuart 1985, Sarkar 1996). Among the reasons for this skepticism, is the fact that the use of that term in biology is not as precise as in the mathematical theory of communication. Second, although the standard account of genetic information refers to an alleged semantic property of genes, it is not clear if and how any genuinely intrinsic semantics is involved.
One possibility for building a theory of information
in biology is to rely on the mathematical theory of communication. The
mathematical theory of communication is a branch of mathematics that arose out
of communication theory. As Shannon and Weaver defined it, “[t]he fundamental
problem of communication is that of reproducing at one point either exactly or
approximately a message selected at another point” (1949: 31). According to Adams (2003: 472), “at the foundation of information theory is the development of methods
to measure the amount of information generated by an event or events, and
mathematical treatments of the transmission characteristics of communication
channels”. It relies on the theory of probability to model information sources,
flow, and communication channels. The amount of information is measured in
terms of the unexpectedness of the sequence of signals, written H=![]() pi
log (1/pi), where pi is the probability of
the ith form of signal.
pi
log (1/pi), where pi is the probability of
the ith form of signal.
This theory allows one to define the amount of information as the measure of the probability of selection of a particular message among the set of all possible messages. The probabilistic measure of information provided by this theory is non-semantic, indifferent to meaning (Shannon & Weaver 1949:31, Cover and Thomas 1999, Jablonka 2002). Despite the fact that the meaning-free concept of information theory can be invaluable in biological research for several purposes (Adami, 2004), controversy continues about a non-semantic understanding of information in biology and whether it is sufficient for a theoretical understanding of information in biology. Jablonka (2002), for instance, argues that this concept is not sufficient by pointing out that, for instance, a DNA sequence encoding a functional enzyme and a same-length sequence coding for a completely non-functional enzyme would contain, according to the above-mentioned measure, the same amount of information. It is obvious, however, that these two messages don’t mean the same thing to the cell. This indicates the necessity of a definition of information in biology which includes a semantic and a pragmatic dimension.
We think that a semantic and pragmatic notion of information and its derivatives grasp some fundamental features of biological systems and processes that might be otherwise neglected. In particular, the concepts of ‘code’, ‘information’, ‘signals’, ‘message’, ‘signaling’, ‘transduction’, and so on must be regarded as necessary for an understanding of the organization of relations in living beings in order to make clear that what happens in such beings is much more than simple chemistry (for details, see Emmeche and Hoffmeyer 1991; Emmeche 1991; El-Hani, Queiroz and Emmeche, forthcoming). For instance, understanding control and regulation in cellular systems without understand cell signaling is not possible. Bray argues that as “about 50% of the genome of a multicellular organism may code for proteins involved in cell signaling, [...] organisms can be viewed as complex information-processing systems, where molecular analysis alone may not be sufficient” (cited by Williams 1997:476-477). Ideker and colleagues (2001:343) consider that one of the consequences of the Human Genome Project has been a strengthening of the view that “biology is an informational science”, and even argue that biological research needs cross-disciplinary scientists which should be educated through teaching of biology as an informational science (p. 365).
The task of building a theory of information in biology becomes more and more important as our knowledge about the structural and functional complexity of living beings increase. In our view, the use of semiotic concepts and theories to interpret information talk can significantly contribute to a precise and coherent formulation of the notion of information in biology. Our aim here is to introduce a model of information as semiosis, grounded on Peircean semiotics. In the next section, we will examine the interrelationships between the concepts of signs, semiosis, and information in Peirce’s formal science of signs.
2. INFORMATION AND SEMIOSIS IN PEIRCE’S SCIENCE OF SIGNS
Peirce’s conception of Semiotics as the ‘formal science of signs’ has had a deep impact in philosophy, psychology, theoretical biology, and cognitive sciences (see Queiroz and Merrell 2005; Freadman 2004; Fetzer 2001; Houser 1997; Deacon 1997; Hoffmeyer 1996; Tiercelin 1995; Freeman 1983). Peirce defined semiosis (meaning process) as an irreducible triadic relation between sign-object-interpretant (S-O-I) (EP 2.171, CP 2.274). That is, according to Peirce, any description of semiosis involves a relation constituted by three irreducibly connected terms, which are its minimal constitutive elements (MS 318:81; CP 2.242):
My definition of a sign is: A Sign is a Cognizable that, on the one hand, is so determined (i.e., specialized, bestimmt) by something other than itself, called its Object, while, on the other hand, it so determines some actual or potential Mind, the determination whereof I term the Interpretant created by the Sign, that that Interpreting Mind is therein determined mediately by the Object” (CP 8.177. Emphasis in the original).
Peirce conceives a ‘Sign’ or ‘Representamen’ as a ‘First’ which stands in such a genuine triadic relation to a ‘Second’, called its ‘Object’, so as to be capable of ‘determining a Third’, called its ‘Interpretant’, to assume the same triadic relation to its Object in which it stands itself to the same Object (CP 2.274. See also CP 2.303, 2.242, 2.92, 1.541). The term ‘Sign’ was used by Peirce to designate the irreducible triadic relation between S, O and I, as well as to refer to the first term of the triad (sometimes ‘Representamen’). Some commentators propose, then, that we should distinguish between ‘Sign in a broad sense’ and ‘Sign in strict sense’ (e.g, Johansen 1993: 62). We will systematically use the term ‘Sign’ in this paper to refer to the first term of the triad, and ‘semiosis’, to refer to the whole triad.
The triadic relation between S, O and I is regarded by Peirce as irreducible, in the sense that it is not decomposable into any simpler relation:
‘... by ‘semiosis’ I mean [...] an action, or influence, which is, or involves, a cooperation of three subjects, such as a sign, its object, and its interpretant, this tri-relative influence not being in any way resolvable into actions between pairs’ (CP 5.484).
Semiosis entails the instantiation of chains of triads. As Savan (1986: 134) argues, an interpretant is both the third term of a given triadic relation and the first term (sign) of a subsequent triadic relation. This is the reason why semiosis cannot be defined as an isolated triad; it necessarily involves chains of triads (see Merrell 1995) (see Figure 1).
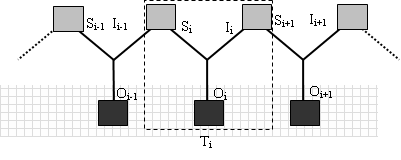
Figure 1: The triadic relation S-O-I forming a chain of triads
Indeed, one of the most remarkable characteristics of Peirce’s theory of signs is its dynamical nature. According to Merrell (1995: 78), “Peirce’s emphasis rests not on content, essence, or substance, but, more properly, on dynamic relations. Events, not things, are highlighted.” The complex (S-O-I) is the focal-factor of a dynamical process (Hausman 1993: 72). As a process thinker, it was quite natural that Peirce conceived semiosis as basically a process in which triads are systematically linked to one another so as to form chains.
Throughout this paper, it is also important to avoid losing from sight the distinction between the interpreter, which is the system which interprets the sign, and the interpretant. The interpreter is described by Peirce as a ‘Quasi-mind’ (CP 4.536), a description which demands, for its proper interpretation, a clear recognition of Peirce’s broad concept of ‘mind’ (Ransdell 1977, Santaella-Braga 1994). It is far from being the case that only conscious beings can be interpreters in a Peircean framework. Rather, a transcription machinery synthesizing RNA from a string of DNA or a membrane receptor recognizing a given hormone, or an ant recognizing a leaf among several other objects in a garden (and so on) can be regarded as interpreters in such a framework. A basic idea in a semiotic understanding of living systems is that these systems are interpreters of signs, i.e., that they are constantly responding to selected signs in their surroundings. In short, the interpreter does not have to be a conscious being, not even an organism, as it may be some part or subsystem within an organism or a humanly-designed product.[2]
We also need to consider here Peirce’s distinctions regarding the nature of objects and interpretants.[3] He distinguishes between the immediate and the dynamical objects of a sign as follows:
“We must distinguish between the Immediate Object – i.e., the Object as represented in the sign – and [...] the Dynamical Object, which, from the nature of things, the Sign cannot express, which it can only indicate and leave the interpreter to find out by collateral experience” (CP 8.314. Emphasis in the original).
Or else:
“... we have to distinguish the Immediate Object, which is the Object as the Sign itself represents it, and whose Being is thus dependent upon the Representation of it in the Sign, from the Dynamical Object, which is the Reality which by some means contrives to determine the Sign to its Representation” (CP 4.536).
And we should also take into account his distinction between the following two kinds of interpretants:[4]
“The Immediate Interpretant is the immediate pertinent possible effect in its unanalyzed primitive entirety. […]. The Dynamical Interpretant is the actual effect produced upon a given interpreter on a given occasion in a given stage of his consideration of the Sign” (MS 339d:546-547. Emphasis in the original).
Let us consider, first, Peirce’s distinction between the immediate and the dynamic objects of a sign. The immediate object of a sign is the object as it is immediately given to the sign, the dynamical object in its semiotically available form. The dynamical object is something which the sign can only indicate, something that the interpreter should find out by collateral experience (EP 2.498; CP 8.178).
In turn, Peirce defines the dynamical interpretant as the actual effect of a sign, while the immediate interpretant is the ‘range of interpretability’ of a sign – the range of possible effects that a sign is able to produce (see Johansen 1993: 166-167). The dynamical interpretant is the instantiation of one of the possible effects established in the immediate interpretant. As the effect of the sign upon the interpreter (or upon some interpreting system), the dynamical interpretant can be treated as being essentially equal to the significance of the sign when seen in a dynamic and process-oriented perspective.
The notions of ‘meaning’, ‘information’, ‘semiosis’ intersect and overlap in different ways (see Johansen 1993). Peirce (see Fitzgerald 1966: 84; Bergman 2000) defined meaning as connected to the triadic relation as a whole (EP 2:429), as well as to different correlates of a triad - e.g., object (MS 11, EP 2:274), interpretant (EP 2:496, EP 2:499; CP 4:536).
For Debrock (1996), Peirce defined ‘information’ at least ordinarily (CP 2.418), metaphysically (CP 2.418), as a connection between form and matter, and logically (W 1.276), as the product of extension and intension of a concept.
Peirce described the function of a sign as that of ‘conveying’ a form from an object to an interpretant (EP 2:391):
“… a Sign may be defined as a Medium for the communication of a Form. [...]. As a medium, the Sign is essentially in a triadic relation, to its Object which determines it, and to its Interpretant which it determines. [...]. That which is communicated from the Object through the Sign to the Interpretant is a Form; that is to say, it is nothing like an existent, but is a power, is the fact that something would happen under certain conditions (EP2, p. 544, n.22).
Accordingly, in the biosemiotic approach to the notion of information we are developing, significantly inspired by Peirce, information is conceived as the communication of a form from O to I through S (figure 2). The communication of a form amounts to the transference of a habit embodied in the object to the interpretant, so as to constrain (in general) the interpretant as a sign or (in biological systems) the interpreter’s behavior.
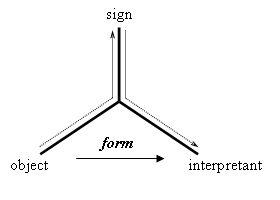
Figure 2: Semiosis as the communication of a form from the object to the interpretant through the mediation of the sign.
Or, to put it in more detailed terms, the production of an effect of the sign on the interpreter results from the communication of the form of the object (as a regularity), via sign, to the interpretant. The interpretant then becomes itself a sign which refers to the object in the same manner in which the original sign refers to it (i.e., there is an invariance in the reconstruction of the form of the object by the interpreter).[5]
According to this approach, ‘information’ can be strongly associated to the concepts of ‘meaning’ and ‘semiosis’. Peirce spoke of signs as ‘conveyers’, as a ‘medium’ (MS 793), as ‘embodying meaning’.
If we consider that Peirce defined a sign both as “a Medium for the communication of a Form” and as “a triadic relation, to its Object which determines it, and to its Interpretant which it determines”, we can say that semiosis is a triadic process of communication of a form from the object to the interpretant by the sign mediation. Therefore, in this framework, we can say that semiosis is information, if we define this latter concept as above.
But what is a Form? Form is defined by Peirce as having the ‘being of predicate’ (EP 2.544) and it is also pragmatically formulated as a ‘conditional proposition’ stating that certain things would happen under specific circumstances (EP 2.388). For Peirce, it is nothing like a ‘thing’ (De Tienne 2003), but something that is embodied in the object (EP 2.544, n. 22) as a habit, a ‘rule of action’ (CP 5.397, CP 2.643), a ‘disposition’ (CP 5.495, CP 2.170), a ‘real potential’ (EP 2.388) or, simply, a ‘permanence of some relation’ (CP 1.415). We can say that Peirce follows a via media in which ‘form’ has both the character of firstness and thirdness. This is in accordance with Bergman’s (2000: 236) proposal of communicated form as a First of a Third.
Form can also be defined as potentiality (‘real potential’, EP 2.388). If we consider this definition, we will also come to the conclusion that form can show the nature of both firstness and thirdness. Consider that potentiality is not the same as mere possibility. For the sake of our arguments, consider Peirce’s treatment of Quality as a ‘mere abstract potentiality’ (CP 1.422). Quality has the nature of firstness, being essentially indeterminate and vague. But we can also talk about a generality of Quality. In this case, we are beyond the domain of pure firstness, as generality refers to some law-like tendency, and thus to the nature of thirdness. Peirce works in this case with a merging of firstness and thirdness. As an abstract potentiality, Quality is closer to a blend of firstness and thirdness than to pure firstness. Such a treatment seems to be compatible with Peirce’s categorical scheme, since, as Potter (1997, p. 94) stresses, “the categoreal structure which Peirce uses is […] highly subtle and complex, admitting of various combinations”.
An understanding of information as a communication of a form or habit embodied in the object to the interpretant through the sign, which brings about a constrained set of effects on the interpreter, can be fruitfully connected to Rosenthal’s (1997) pragmatic approach to meaning, but with some qualification. Rosenthal argues that meaning is an emergent relational pattern of behavior. Information, when conceived as the communication of a form from O to I through the mediation of S, can be seen as a process working as a constraining factor of possible patterns of interpretative behaviors or processes. As meaning is also defined by Peirce as something communicated in semiosis (NEM 4:309), we will opt here for explaining meaning as being associated with the interpretant, which, after all, embodies the reconstructed form of the object.
Peirce (CP 8.177) writes that a sign determines an interpretant in some ‘actual’ or ‘potential’ Mind (in other passages, a ‘quasi-mind’. See CP 4.536). We take this suggestion to introduce in our analysis a differentiation between ‘potential’ and ‘effective’ semiosis. Potential semiosis is defined as a triadically-structured process which is not taking place, which is only in potency. Effective semiosis, in turn, is a sign in effective action, i.e., a sign which, by being actualized, has an actual effect on the interpreter.
Following the distinction between potential and effective semiosis, we can define potential information as a process of communicating a form which could be realized in a given moment, while effective information is the communication of a form from an object to an interpretant through the sign, i.e., a sign in effective action.
The notion of information as form communicated from O to I through the mediation of S allows us to conceive it in a processual way, as a constraining factor of possible patterns of interpretative behavior. When applying this general semiotic approach to biological systems, information will most often be an interpreter-dependent objective process. It cannot be dissociated from the notion of a situated (and actively distributed) communicational agent (potential or effective). It is interpreter-dependent in the sense that information triadically connects representation (sign), object, and an effect (interpretant) on the interpreter (which can be an organism or a part of an organism). In turn, the habit or form which is communicated in information is embodied in the object, treated in the Peircean framework as (dynamically) the primary constraining factor of interpretative behavior.[6] Thus, the form - as a regularity in the object – acts as constraint on the interpreter’s behavior, but the interpreter always reconstructs the form of the object when interpreting a sign. Nevertheless, the interpreter does so in such a manner that an invariance is retained, which makes it possible, in fact, the very act of interpretation.
In sum, according to our interpretation of Peirce’s remarks quoted above, information has a processual nature: information is the process of communicating a form from the object to the interpretant through the sign. A framework for thinking about information as a process can be built in Peircean terms by employing the following definitions:
[Information = semiosis] A triadic-dependent process through which a form embodied in the object in a regular way is communicated to an interpretant through the mediation of a sign.
[Potential information = potential semiosis] A process of communicating a form from an object to an interpretant through the mediation of a sign that could take place in a given moment, changing the state of the interpreter.
[Effective information = effective semiosis] The process by which a sign effectively exerts an effect (interpretant) on some system (an interpreter) by making the interpretant stands in a similar relation to something else (the object of the sign) as that to which the sign stands, thus mediating the relation between object and interpretant. The sign effectively communicates, thus, a form from the object to the interpretant, changing the state of the interpreter.
To formulate the above definitions in a sufficiently clear way, we should define what we mean by ‘process’. We follow here Rescher in his definition of a process as “... a coordinated group of changes in the complexion of reality, an organized family of occurrences that are systematically linked to one another either causally or functionally” (Rescher 1996:38).
These definitions certainly raise several questions and face a number of difficulties when they are seen against the background of other accounts of information. We shall leave to a subsequent paper, however, a discussion about how they relate to and result in controversies concerning a number of ideas about what is information expressed by different authors.
3. A BIOLOGICAL EXAMPLE: A SEMIOTIC ANALYSIS OF THE GENETIC INFORMATION SYSTEM
As we argued above, the concept of information has been playing an important role in the biological sciences since the mid-20th century, but it remains a problematical notion, to which biologists have ascribed a metaphorical role. Understandably, it is a controversial issue whether the concept of information and its derivatives should be maintained or eliminated from biology. We assume that they should be kept in the biological conceptual framework, but a semantic/pragmatic theory of information in biology should be built, in order to ascribe precise meanings to those concepts and to articulate them in proper explanations of sign processes in biological systems. As a way of contributing to this endeavor, we have been using semiotic concepts and theories to build an account of genes as signs and genetic information as semiosis grounded on Peircean semiotics (El-Hani, Queiroz and Emmeche, forthcoming). As an example of our application of the conceptual tools introduced in the previous section, we will present here part of the results we obtained so far.[7]
When applying Peirce’s semiotics to understand the nature of genetic information, it is inevitable to engage in interpretation about how to see, for instance, the relationship between what molecular biologists and biophysicists call forms of information processing (i.e., production and interpretation of signs) in a complex living system such as the cell and forms of causality in that system. So, the analysis of the genetic information system given below is not the only way to apply Peircean semiotics to this particular case; and some might object to the particular way we addressed the problem. We also acknowledge that there are peculiar features of the genetic information system which do not exactly conform to any standard Peircian framework. Nevertheless, we think that we have been faithful both to the basic insights and concepts of semiotics and to the findings of molecular biology, and that the few changes we have made in specific semiotic conceptions (as we shall explicate below) are necessitated by the growth of scientific knowledge about the system analyzed.
3.1. SOME BASIC NOTIONS ABOUT THE GENETIC INFORMATION SYSTEM
First, we should briefly introduce some basic notions about the genetic information system.[8] Let us consider a very simple model of the process of gene expression (Figure 3). During the synthesis of pre-mRNA (transcription), the four-base language of DNA (as a sequence of nucleotides including the bases adenine, A, guanine, G, cytosine, C, and thymine, T) is copied or ‘transcribed’ into the four-base language of RNA (with uracil, U, replacing T).
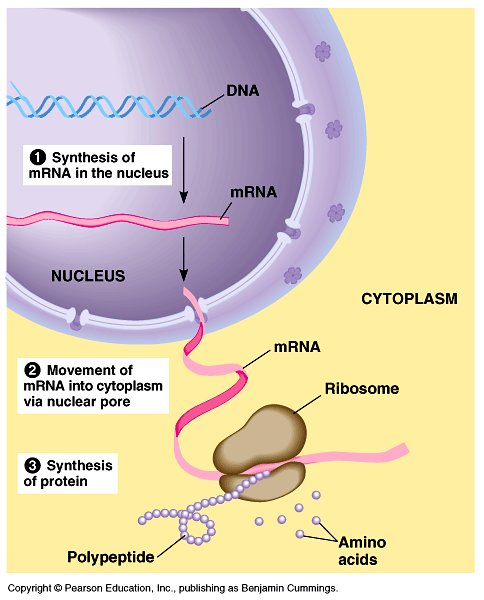
Figure 3: A model of the steps of gene expression (From Campbell & Reece, 2002). Many details are omitted, for the sake of simplicity.
The effects of a protein-coding gene on a given cell or organism are regulated mainly by control of gene expression at the level of transcription initiation. The transcription of a gene can be either repressed, when the corresponding mRNA and encoded protein or proteins are synthesized at low rates or not synthesized at all, or activated, when both the mRNA and encoded protein or proteins are, ceteris paribus, produced at much higher rates. Through the control of gene expression, only a subset of all genes present in any cell type in a multicellular organism is really expressed. Thus, from all the potential protein products a given cell type might exhibit, only a specific number and variety will be present. This is the fundamental basis for cell differentiation in multicellular organisms.
In the end of the 1970s, it was found that eukaryotic genes are split into pieces of coding sequence, named ‘exons’, separated by non-coding segments, named ‘introns’ (after Gilbert 1978). The vast majority of genes in multicellular eukaryotes contain multiple introns and the presence of such introns allows for the expression of multiple related proteins from a single stretch of DNA by means of a process known as ‘alternative splicing’ (see below).
In eukaryotic protein-coding genes, introns are excised from a long ‘primary transcript’ (precursor mRNA or pre-mRNA), i.e., the RNA copy of an entire DNA sequence containing both exons and introns, in a process known as RNA ‘processing’, which includes other events not described here. After the introns are excised, the coding exons are joined back together into a functional mRNA, which will be transported to the cytoplasm of the eukaryotic cell, where protein synthesis will take place.
The effects of genes on the functioning of a cell or organism can also be regulated by means of alternative pre-mRNA splicing, so as to produce different gene products from the same pre-mRNA. Alternative RNA splicing is an important mechanism for the production of different forms of proteins (isoforms) by different cell types. The fibronectin (FN) gene, for instance, generates more than 20 different FN isoforms. The FN gene has approximately 75,000 nucleotides (75-Kb) and contains numerous exons. After the FN pre-mRNA is transcribed from DNA, it undergoes cell type-, development- and age-specific splicing. Each FN isoform is encoded by a differently, alternatively spliced mRNA, and, therefore, each isoform results from a unique combination of exons found in the FN gene (see Figure 4).
Consider, for instance, the splicing of FN pre-mRNA in fibroblasts and hepatocytes. In fibroblasts, splicing of the FN pre-mRNA results in mRNAs containing exons EIIIA and EIIIB. The fibroblast FN isoform contains amino acid sequences that bind tightly to proteins in the plasma membrane. This specific FN isoform contributes to the adhesion of fibroblasts to the extracellular matrix. In hepatocytes, the major cell type in the liver, cell-type specific splicing results in functional FN mRNAs lacking exons EIIIA and EIIIB. FN secreted by hepatocytes does not adhere tightly to fibroblasts or most other cell types, freely circulating in the blood stream. When the wall of a vase is ruptured, hepatocyte FN plays a fundamental role in the formation of blood clots, due to the presence in the protein of fibrin-binding domains, amino acid sequences that bind to fibrin, one of the main constituents of blood clots. When hepatocyte FN is bound to fibrin, it interacts with integrins, cell-adhesion protein molecules found in the membranes of activated platelets. As a result, the blood clot is expanded through the addition of platelets.
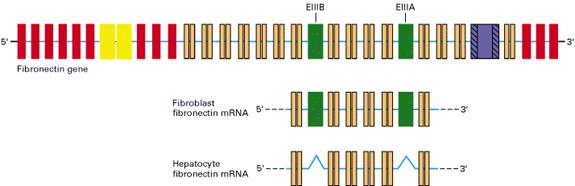
Figure 4: Cell type-specific splicing of fibronectin pre-mRNA in fibroblasts and hepatocytes. The 75-kb FN gene (top) contains multiple exons. Introns are shown in the diagram as thin lines and are not drawn to scale. Most of the introns are much longer than any of the exons. The FN mRNA produced in fibroblasts includes the EIIIA and EIIIB exons, whereas these exons are spliced out of FN mRNA in hepatocytes (from Lodish et al. 2000. Available at http://www.ncbi.nlm.nih.gov/PubMed/).
Finally, translation is an essential part of protein synthesis, consisting in the process by which the nucleotide sequence of an mRNA serves as a template for the synthesis of a polypeptide chain, i.e., for a series of events in which amino acids are ordered and joined to form the primary structure of a protein. Three types of RNA molecules are involved in translation, performing different but cooperative functions. mRNAs are the ‘vehicles’ of the genetic information transcribed from DNA. The ‘message’ at stake is ‘written’ in the form of a series of three-nucleotide sequences, called ‘codons’, each of which specifying a particular amino acid. tRNAs play a fundamental role in the process of deciphering the codons in mRNA. Each type of amino acid has its own subset of tRNAs. They act as transporters, binding amino acids and carrying them to the growing end of a polypeptide chain in response to specific codons in the mRNA. The reason why the correct tRNA with its attached amino acid is selected at each step in protein synthesis lies in the fact that each specific tRNA molecule contains a three-nucleotide sequence, called an ‘anticodon’, that base-pairs with its complementary codon in the mRNA. In this manner, for each specific codon in mRNA a specific amino acid, carried by a specific tRNA, is included in a polypeptide chain, according to the rules expressed in the almost universal ‘genetic code’. Along with 100 different proteins, several types of rRNA are components of ribosomes, the complex and large macromolecular structures that act, so as to say, as guides to coordinate the assembly of the amino acid chain of a protein. In fact, a rRNAs (a ribozyme), and not a protein, is probably the catalyst involved in the formation of peptide bonds in protein synthesis.
Recognition of a codon in mRNA specifying a given amino acid by a particular tRNA is, in fact, the second step in ‘decoding’ the genetic ‘message’. The first step is the attachment of the appropriate amino acid to a tRNA in a reaction catalyzed by a specific aminoacyl-tRNA synthetase. The specificity of the attachment between amino acids and tRNAs results from the capacity of each one of these enzymes of recognizing one amino acid and all its compatible, or ‘cognate’, tRNAs. Therefore, the rules captured in the genetic code ultimately depend on the recognition activity of aminoacyl-tRNA synthetases.
Although the terms ‘translation’ and ‘protein synthesis’ are usually employed interchangeably, this is not correct, since, although translation is obviously an essential step in protein synthesis, this process involves further steps. Polypeptide chains undergo post-translational folding and often other changes, as, for instance, chemical modifications and association with other polypeptide chains, that are required for production of functional proteins. All these steps in protein synthesis can undergo regulation.
3.2. A SEMIOTIC ANALYSIS OF GENES AND GENETIC INFORMATION
If we take Peirce’s concepts of sign and semiosis as bases for analyzing what is a gene, it will be the case that the action of a gene as a sign will have to be understood as a relationship between three elements (Figure 5). Given the account of information developed in this paper, genetic information can be described as a semiotic process. In these terms, there’s more to genetic information than just sequences of nucleotides in DNA.

Figure 5: A semiotic analysis of the gene as a sign
In this picture, a string of DNA (say, the FN gene) is treated as a sign. As a protein-coding gene, the FN gene, for instance, stands – in a triadic-dependent relation – for a specific sequence of amino acids (immediate object) – one of the FN isoforms, translated out of a mature mRNA after alternative splicing (which, as the figure shows, can take place or not, depending on the string of DNA we are analyzing)[9] – through a process of reconstruction of a specific form (interpretant).[10]
Given that a sign is the mediating element in a semiotic process through which a form is communicated from an object to an interpretant, we treat the interpretant here as the reconstruction of a form (habit) which was embodied in an object. To be more explicit, we defined above information as the communication of a form from the object to the interpretant, and we also argued that such a communication will in turn constrain the behavior of the interpreter. What we mean by ‘reconstruction’ here is a process by which the form of a protein in a cell generation is communicated through signs in DNA (in potency) to a cell in a subsequent generation. Thus, a regularity obtains (with possible evolutionary consequences) in the three-dimensional structure and function of proteins along generations.
We will introduce the qualifiers ‘composite’ and ‘simple’ to incorporate a part-whole relationship in our semiotic analysis of genes, referring to a stretch of DNA or mature RNA as a whole as a composite sign, formed by clusters of simple signs, codons. We can now introduce in our analysis the distinction between immediate and dynamical object, and immediate and dynamical interpretant.
In a Peircean framework, the immediate object can be understood as the characteristics selected in the sign as a means of indicating the dynamical object. It is not the case, in this framework, that the immediate object is a condition of possibility for the dynamical object. Nevertheless, in the case we are analyzing here the interpreter creates a dynamical object of a given class (showing a given habit) on the grounds of indications present in the sign.
A cell uses signs in DNA as a basis for synthesizing a dynamical object sufficiently resembling a past dynamical object which does not exist anymore but resulted in successful, adaptive experiences. This is the reason why we claim that, in this case, the immediate object establishes conditions of possibility for the dynamical object.
The dynamical object of a gene is a functional, folded, and chemically modified protein, which is often not entirely specified in the sequences of nucleotides or amino acids, but it is rather indicated by such sequences.[11] Functional proteins are not always simply translated out of nucleotide sequences by a cell, but they are rather found out through resources the cell acquire by collateral experience, i.e., by habits that a cell acquire in its development towards the states characteristic of a given cell type, and can be traced back to evolutionary processes.[12] A functional FN isoform, for instance, is a dynamical object.
The composite immediate object of a protein-coding gene is the sequence of amino acids of a polypeptide, as this is the object represented in the gene’s vehicle, a string of DNA. Each amino acid, in turn, is a simple immediate object. If we consider the sequence of amino acids of a specific FN isoform, we will say, in the terms of our analysis, that such a sequence is an immediate object of the FN gene. It is important to bear in mind, however, that it is an immediate object, not the immediate object. After all, the FN gene codes more than 20 different FN isoforms, all of them being possible immediate objects of the FN gene as a sign in DNA.
The sequence of amino acids, the composite immediate object, is the dynamical object in its semiotically available form. The sequence of amino acids of each FN isoform amounts to a specific protein coded – in its semiotically available form – in a mature RNA which results, after splicing, from a pre-mRNA transcribed from the FN gene.
The immediate object, a sequence of amino acids, can indicate a range of possible functional proteins, dynamical objects, as a single amino acid sequence can be folded in different ways in different cellular contexts. But we should not lose from sight, however, that such an indication by the immediate object plays a fundamental role in the reconstruction of the dynamical object, since it is not the case that any three-dimensional protein can be produced from a given amino acid sequence.[13]
The immediate interpretant of a codon as a simple sign is the range of interpretability established by the rules of base pairing by which specific nucleotides in DNA determine specific nucleotides in mRNA, or the range of interpretability of three-nucleotide sequences in mature mRNA as established in the genetic code, a set of rules by means of which nucleotide sequences determine the addition of specific amino acids to a growing polypeptide chain (Figure 6).[14] The dynamical interpretant of a codon as a simple sign amounts, then, to the realization of one of the rules of base pairing or of the genetic code.
A composite sign in DNA determines a range of possible composite immediate objects. It is true that there are cases in which a stretch of DNA codes for only one protein product. In this case, the composite sign in DNA determines only one immediate object. Nevertheless, in eukaryotic cells at least, most stretches of DNA codes for several distinct proteins, as in the case of the FN gene. Therefore, we can define the immediate interpretant of a composite sign as the range of interpretability of that sign in DNA, i.e., as the possible immediate objects, the possible sequences of amino acids, that can be produced from that sign in DNA. Alternative RNA splicing is understood, in these terms, as one of the processes that enrich the range of interpretability, the immediate interpretant, of a stretch of DNA. In the case of the FN gene, its immediate interpretant comprises more than 20 possible composite immediate objects.

Figure 6: The genetic code. Sets of three nucleotides (codons) in an mRNA molecule are translated into amino acids during protein synthesis according to the rules shown in the table above. (from Griffiths et al. 1999. Available at http://www.ncbi.nlm.nih.gov/PubMed/).
This analysis is in accordance with the definition of a sign as medium for communicating the form of an object to an interpretant. The interpretant can be seen, thus, as a reconstruction of the form of an object. It follows that the immediate interpretant of a stretch of DNA or mRNA as a composite sign, i.e., its range of interpretability, amounts to the diversity of possibilities of reconstruction of the form of the composite immediate object, the sequence of amino acids in a polypeptide.
The dynamical interpretant of a stretch of DNA or mRNA as a composite sign corresponds to the effective reconstruction of a sequence of amino acids. In an alternatively spliced gene, such as the FN gene, this realization involves the instantiation of a specific splicing pattern in a given cell type, at a given developmental stage. Thus, one of the possibilities established in the range of interpretability of a stretch of DNA, in its immediate interpretant, is actualized. In a fibroblast, for instance, when a specific immediate object is synthesized, the fibroblast-specific FN isoform, this means that, from the range of possible sequences of amino acids that might be made out of the FN gene – its immediate interpretant –, a specific sequence was reconstructed – its dynamical interpretant.
After it is actualized, an immediate object indicates a particular dynamical object – say, a specific FN isoform. It is the dynamical object, then, that has an effect on the cell as a global interpreter. We can define, then, a dynamical interpretant of the dynamical object, a particular effect on a cell, among a range of possible effects – the immediate interpretant of the dynamical object. This dynamical interpretant is the actualization of one of the possible effects that a composite sign might have on the interpreter. Its range of interpretability is the immediate interpretant of the composite sign.
This analysis faces the potential problem that it seems to treat the sign as the primary constraining factor in semiosis, while this role is reserved for the dynamical object in Peirce’s theory of signs. After all, we are describing here how S (a sequence of nucleotides in DNA) determines O (a sequence of amino acids in a polypeptide) through I (a range of possibilities of reconstruction of sequences of amino acids).[15] We accommodate this description into a Peircean framework by examining the constraining action of the object in evolutionary terms (see Figure 7). Consider two different generations of a population, in times t1 and t2, and a protein (dynamical object) in t1 that increases the likelihood of successful, adaptive experiences of organisms possessing it. Therefore, that protein increases the likelihood that a gene (sign) encoding it will be present in high frequencies in the next generation, in t2. Indeed, the sequence of a gene is determined, by past natural selection, because of the effects it produces (Maynard Smith 2000:177). This gene, in turn, will bring to the next generation the potency to produce that protein, as a dynamical object, by indicating it through its semiotically available form, its immediate object. Signs in DNA will carry to future generations the potentiality of reconstructing the form of that protein in generations to come. This means that that gene, as a sign, exerts a determining influence on the range of possibilities of reconstructing sequences of amino acids in the next generation. If we follow this set of ideas, we will be able to see how, in evolutionary terms, O determines I through S, in conformity with Peirce’s account of semiosis. Nevertheless, the role of O as the primary constraining factor of semiosis depends, in the genetic information system, on the role of S, in a given generation, in determining O through I. We can say, in short, that the fact that S determines O through I in a given population in t2 is itself determined by the fact that O determined I by increasing the likelihood of S being present in a high frequency in t2, by means of its involvement in successful experiences in t1.
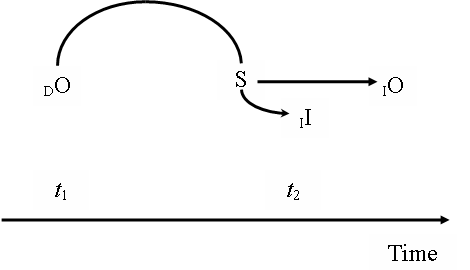
Figure 7: The dynamical object (functional protein) as the primary constraining factor of semiosis in the genetic information system. S, sign; DO, dynamical object, IO, immediate object; II, immediate interpretant; t, generation time.
The relationship between signs in DNA and the sequence of amino acids of a protein (the composite immediate object) is established by a complex mechanism of interpretation, involving transcription, RNA processing and translation. Thus, to interpret a string of DNA, more than one interpretative system is required, including, for instance, RNA polymerases, involved in the transcription of DNA into RNA, and ribosomes, involved in the translation of mRNA into proteins. These interpretative systems are parts or subsystems of a cell as a global interpreter, and their actions are subordinated to the latter. That ultimately the whole cell participates in the network necessary for the interpretation that is demanded for the effect of a gene product to take place (cf. Emmeche & Hoffmeyer 1991) is shown by the impressive array of signaling pathways regulating the interpretation of Signs in DNA.
Symptomatically, a Peircean approach to the gene concept entails that genetic structures should not be seen in isolation from the larger system by which they are interpreted. From this perspective, the meaning of a gene to its interpreter, the cell, or, to put it differently, the biological meaningfulness of a gene, is found not only in DNA sequences in a chromosome. After all, according to this approach, there is more to genetic information than just a sequence of nucleotides in DNA. We will have to include the effect of the gene-as-a-sign on the cell or organism, and, in fact, the very role of cellular subsystems as interpreters of strings of DNA, in such a way that they relate signs to specific dynamical objects, proteins which play a function inside the cellular system and have an effect on it or on the organism of which the cell is a part.
In sum, the semiotic analysis of the genetic information system developed above leads to the following conclusions:
(i)i.
Genes should be regarded as signs
in DNA, which can only have any effect on a cell through a triadic-dependent
process (semiosis);
(ii)ii.
This process is genetic
information and involves more than just genes as signs in DNA but also objects
and interpretants;
(iii)iii.
Genetic information is the process
by means of which a form in a dynamical object (a functional protein) is
communicated to an interpretant (the reconstruction of a specific sequence of
amino acids in a cell) by means of signs in DNA. [16]
4. CONCLUDING REMARKS
There is controversy about the prospects of the non-semantic understanding of information offered by the mathematical theory of communication for developing a theory of information in biology. Despite the usefulness of this theory for several purposes in biological research, there is still doubt whether it is sufficient for the understanding of biological information. We believe that biology needs a semantic/pragmatic account of information. Throughout this paper, we argue that semiotic concepts and theories, and, in particular, C. S. Peirce’s formal science of signs, can lead to a precise and coherent formulation of the notion of information in biology.
Toward this end we have developed a model of information as semiosis, grounded on Peircean semiotics. According to this model, information is conceived as the communication of a form or habit from O to I through S, so as to constrain (in general) the meaning of the interpretant as a sign or (in biological systems) the interpreter’s behavior. Or, to put it differently, information is the same as semiosis, i.e., a triadic-dependent process through which a form embodied in the object in a regular way is communicated to an interpretant through the mediation of a sign.
The treatment of information in biology based on Peircean semiotics leads to an account of genes as signs and genetic information as semiosis. According to this account, genetic information is a triadic relation between a (composite) sign in DNA, i.e., the sequence of nucleotides of a gene; an immediate object, i.e., the sequence of amino acids of a polypeptide (or the sequence of nucleotides of a RNA); and an immediate interpretant, i.e., the possibilities of reconstruction of specific immediate objects, which amount to the range of interpretability of a (composite) sign in DNA. Genetic information has, thus, a processual nature and should not be identified with sequence information in a nucleic acid. Rather, it is a process by means of which a form in a dynamical object (a functional protein) is communicated to an interpretant (the reconstruction of a specific sequence of amino acids in a cell) by means of signs in DNA.
ACKNOWLEDGEMENTS
J.Q. thanks The State of São Paulo Research Foundation (FAPESP) for the post-doctoral studies grant n. 02/09763-2.
C.E. thanks the Faculty of Science, University of Copenhagen. C.N.E.H. thanks the Brazilian National Research Council (CNPq) for research grants n. 302495/02-9 and 301832/04-8, post-doctoral studies grant n. 200402/03-0, and funding of the project n. 402708/2003-2.
We are thankful to Edwina Taborsky and Peter Harries-Jones for sending us written comments on a preliminary version of the paper. We are also thankful to Wolfgang Hofkirchner and Toshiyuki Nakajima for their comments about the paper.
REFERENCES
Adami, C. 2004. Information theory in molecular biology. Physics of Life Reviews 1:3-22.
Adams, F. 2003. The Informational Turn in Philosophy. Minds and Machines 13: 471–501.
Alberts, B.; Johnson, A.; Lewis, J.; Raff, M.; Roberts, K.and Walter, P. 2002. Molecular Biology of the Cell. 4th Ed. New York: Garland Science.
Bergman, M. 2000. Reflections on the Role of the Communicative Sign In Semeiotic. Transactions of the Charles S. Peirce Society: A Quarterly Journal in American Philosophy. Spring, XXXVI (2): 225-254.
Brunning, J. 1997. Genuine triads and teridentity. In N.Houser, D.Roberts, J.Evra. Eds. Studies in the logic of Charles Sanders Peirce. Indiana: Indiana University Press. pp. 252-270.
Campbell, N. A. and J. B.Reece. 2002. Biology 6th Ed. San Francisco: The Benjamim/Cummings Publ. Co.
Cooper, G. M. and R. E.Haussman. 2003. The Cell: A Molecular Approach. 3rd Ed. Sunderland-MA: Sinauer.
Cover, T.M. and J.A.Thomas. 1999 Information Theory. In R. A. Wilson and F.C. Keil. Eds., MIT Encyclopedia of the Cognitive Sciences. Pp. 404-406. Cambridge-MA: MIT Press.
Deacon, T. 1997. The Symbolic Species: The Co-evolution of Language and the Brain. New York: W.W. Norton & Company.
Debrock, G. 1996. Information and the metaphysical status of the sign. In V. Colapietro & T. Olshewsky Eds. Peirce’s Doctrine of Signs – theory, applications, and connections. Pp.80-89. Berlin, New York: Mouton de Gruyter.
De Tienne, A. 2003. Learning qua semiosis. S.E.E.D. (3): 37-53.
El-Hani, C.N.; Queiroz, J. & Emmeche, C. (Forthcoming). A Semiotic Analysis of the Genetic Information System. Semiotica.
Emmeche, C. 1991. A semiotical reflection on biology, living signs and Artificial Life. Biology and Philosophy 6:325-340.
____. Causal processes, semiosis, and consciousness. 2003. In J. Seibt Ed. Process Theories: Crossdisciplinary Studies in Dynamic Categories, 313-336. Dordrecht: Kluwer.
Emmeche, C. and J. Hoffmeyer. 1991. From language to nature - the semiotic metaphor in biology. Semiotica 84(1/2):1-42.
Fetzer, J. 2001. Computers and Cognition: Why Minds are not Machines. Dordrecht: Kluwer Academic Publishers.
Fitzgerald, J. 1966. Peirce’s Theory of Signs as Foundation for Pragmatism. University of Notre Dame: Mouton & Co.
Freadman, A. 2004. The Machinery of Talk – Charles Peirce and the Sign Hypothesis. Stanford: Stanford University Press.
Freeman, E. 1983. The Relevance of Charles Peirce. La Salle, Illinois: Monist Library of Philosophy.
Gilbert, W. 1978. Why genes in pieces? Nature 271:501.
Godfrey-Smith, P. 1999. Genes and codes: Lessons from the philosophy of mind? In V.G. Hardcastle Ed. Biology Meets Psychology: Philosophical Essays. Cambridge-MA: MIT Press.
____. 2000. Information, arbitrariness, and selection: Comments on Maynard Smith. Philosophy of Science 67(2), 202-207.
Griffiths, A. J. F.; Gelbart, W. M.; Miller, J. H. and Lewontin, R. C. 1999. Modern Genetic Analysis. New York: W. H. Freeman & Co.
Griffiths, P. 2001. Genetic information: A metaphor in search of a theory. Philosophy of Science 68(3), 394-403.
Hausman, C. 1993. Charles Sanders Peirce’s Evolutionary Philosophy. Cambridge University Press.
Hoffmeyer, J. 1996. Signs of Meaning in the Universe. Bloomington and Indianapolis: Indiana University Press.
Houser, N. 1997. Introduction: Peirce as a logician. In N. Houser, D. Roberts and J. Evra Eds. Studies in the Logic of Charles Sanders Peirce. Pp. 1-22. Indiana: Indiana University Press.
Ideker, T.; T. Galitski, and L. Hood. 2001. A new approach to decoding life: Systems biology. Annual Review of Genomics and Human Genetics 2:343-372.
Jablonka, E. 1994. Inheritance systems and the evolution of new levels of individuality. Journal of Theoretical Biology 170:301-309.
____. 2002. Information: Its interpretation, its inheritance, and its sharing. Philosophy of Science 69: 578-605.
Jablonka, E. and E. Szathmáry. 1995. The evolution of information storage and heredity. Trends in Ecology and Evolution 10:206-211.
Jablonka, E.; M.J. Lamb, and E. Avital. 1998. ‘Lamarckian’ mechanisms in Darwinian evolution. Trends in Ecology and Evolution 13:206-210.
Johansen, J. 1993. Dialogic Semiosis. Indiana: Indiana University Press.
Lewin, B. 2000. Genes VII. Oxford: Oxford University Press.
Lizska, J.J. 1990. Peirce’s interpretant. Transactions of the Charles S. Peirce Society, Summer Vol. XXVI (1): 17-61.
Lodish, H.; Berk, A.; Matsudaira, P.; Kaiser, C. A.; Krieger, M.; Scott, M. P.; Zipursky, S. L. and Darnell, J. 2003. Molecular Cell Biology . 5th Ed. New York: W. H. Freeman & Co.
Maynard Smith, J. 2000.The concept of information in Biology, Philosophy of Science 67(2): 177-194.
Maynard Smith, J. and E.Szathmáry. 1995. The Major Transitions in Evolution. Oxford: W. H. Freeman.
__. 1999. The Origins of Life: From the Birth of Life to the Origins of Language. Oxford: Oxford University Press.
Merrell, F. 1995. Peirce’s Semiotics Now. Toronto: Canadian Scholar’s Press.
Murphey, M. 1993. The Development of Peirce’s Philosophy. Cambridge: Harvard University Press.
Nöth, Winfried 1995. Handbook of Semiotics. Bloomington and Indianapolis: Indiana University Press.
Oyama, S. 2000. The Ontogeny of Information: Developmental Systems and Evolution. 2nd Ed.. Cambridge: Cambridge University Press.
Parker, K. 1998. The Continuity of Peirce’s Thought. Nashville:Vanderbilt University Press.
Peirce, Charles Sanders. 1931–1935. The Collected Papers of Charles Sanders Peirce. [C. Hartshorne and P. Weiss Eds. Cambridge-MA: Harvard University Press, 1931–1935], Vols. VII–VIII [A. W. Burks Ed. same publisher, 1958]. Electronic edition reproducing Vols. I–VI. Charlottesville: Intelex Corporation. (Here referred as CP, followed by volume and paragraph number.)
____. 1967. Annotated Catalogue of the papers of Charles S. Peirce. Amherst-MS: University of Massachusetts. Robin, R. Ed. [References to manuscripts and letters by Charles S. Peirce — MS and L — are in accordance with this catalogue.]
____. [1893-1913] 1998. The Essential Peirce: Selected Philosophical Writings. Vol. II. (Ed.) Peirce Edition Project. Bloomington and Indianapolis: Indiana University Press. (Here referred as EP2, followed by the number of the page.)
____. [1839-1914] 1982-2000. Writings of Charles S. Peirce: a Chronological Edition. Vol. 2. Ed. Peirce Edition Project. Bloomington: Indiana University. [Quoted as W, followed by page number].
____. 1976. New Elements of Mathematics by Charles S. Peirce, Carolyn Eisele Ed. The Hague: Mouton. [Quoted as NEM, followed by page number].
Potter, V. 1997. Charles S. Peirce: On Norms and Ideals. University of Massachusetts Press.
Queiroz, J. and F. Merrell, F. 2005. Abduction – between subjectivity and objectivity. Semiotica (Special Issue on Abduction) 153 (1/4).
Ransdell, J. 1977. Some leadings ideas of Peirce’s semiotic. Semiotica 19(3/4): 157-178.
Rescher, N. 1996. Process Metaphysics: An Introduction to Process Philosophy. New York: State University of New York Press.
Rosenthal, S. 1997 Pragmatic Experimentalism and the Derivation of the Categories, In. J. Brunning and P. Forster. Eds. The Rule of Reason. pp. 120-138 Toronto: University of Toronto Press..
Santaella-Braga, L. 1994. Peirce’s broad concept of mind. European Journal for Semiotic Studies 6(3,4):399-411.
Sarkar, S. 1996. Biological information: A skeptical look at some central dogmas of molecular biology. In S. Sarkar Ed. The Philosophy and History of Molecular Biology: New Perspectives. Dordrecht: Kluwer.
__. 2000. Information in genetics and developmental biology: Comments on Maynard Smith, Philosophy of Science 67(2), 208-213.
Savan, D. 1986. Response to T. L. Short. Transactions of the Charles S. Peirce Society: A Quarterly Journal in American Philosophy. Summer, XXII (2), 125-143.
__. 1987-1988. An Introduction to C. S. Peirce’s Full System of Semeiotic. Toronto Semiotic Circle. Monograph Series of the TSC, Number 1.
Shannon, C. E. and W. Weaver. 1949. The Mathematical Theory of Communication. Urbana-IL: University of Illinois Press.
Short, T. L. 1996. Interpreting Peirce’s interpretant: a response to Lalor, Liszka, and Meyers. Transactions of the Charles S. Peirce Society. Fall Vol. XXXII (4): 488-541.
___. 1998. What's the use? Semiotica 122(1/2): 1-68.
Sterelny, K. 2000. The ‘genetic program’ program: A commentary on Maynard Smith on information in Biology. Philosophy of Science 67(2), 195-201.
Stuart, C. I. J. M. 1985. Bio-informational equivalence. Journal of Theoretical Biology 113, 611-636.
Tiercelin, C. 1995. The relevance of Peirce’s semiotic for contemporary issues in cognitive science. In L.Haaparanta and S.Heinamaa Eds. Mind and Cognition: Philosophical Perspectives on Cognitive Science and Artificial Intelligence. Pp. 37-74. Acta Philosophica Fennica 58.
Williams, N. 1997. Biologists cut reductionist approach down to size. Science 277 (5325), 476-477.
Wynnie, J. A. 2000. Information and structure in Molecular Biology: Comments on Maynard Smith. Philosophy of Science 67(3), 517-526.